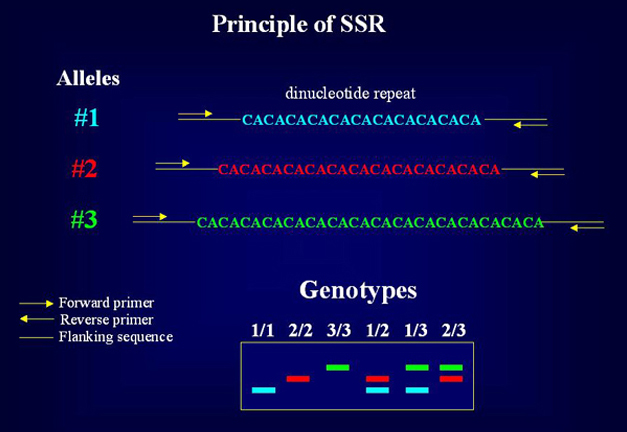
SciELO - Brasil - Microsatellite markers: what they mean and why they are so useful Microsatellite markers: what they mean and why they are so useful

Comprehensive functional analysis and mapping of SSR markers in the chickpea genome (Cicer arietinum L.) - ScienceDirect

Development of Genome-wide SSR Markers for Physical Map Construction with PCR-based Polymorphic SSRs in Jute (Corchorus Spp.) | SpringerLink

Diversity | Free Full-Text | SSR Markers for Trichoderma virens: Their Evaluation and Application to Identify and Quantify Root-Endophytic Strains
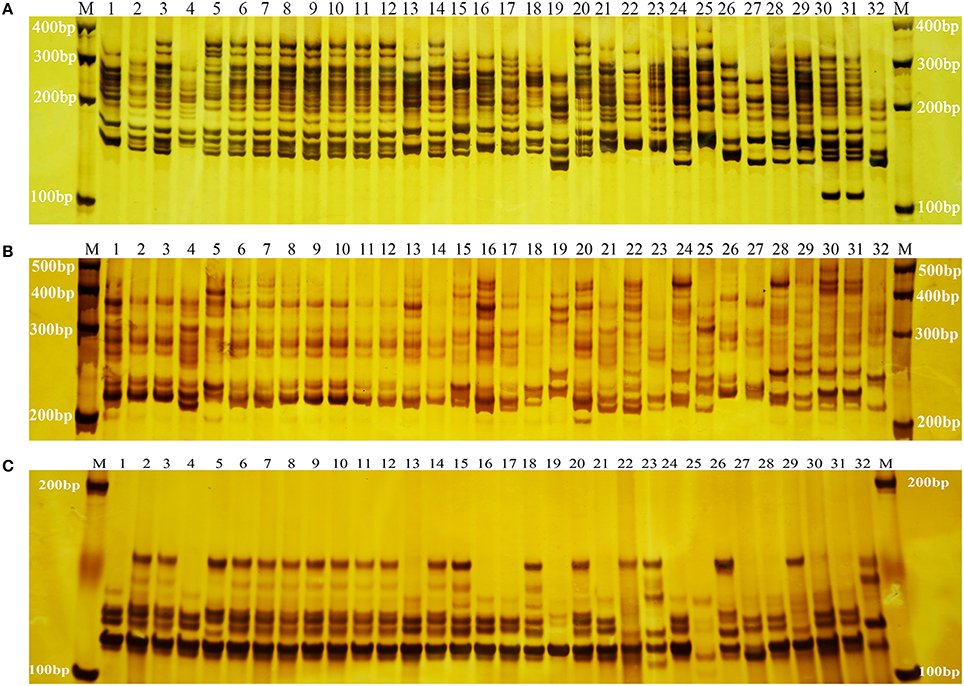
Frontiers | Development of SSR Markers and Assessment of Genetic Diversity in Medicinal Chrysanthemum morifolium Cultivars

EST-SSR markers derived from an elite barley cultivar (Hordeum vulgare L. 'Morex'): polymorphism and genetic marker potential. | Semantic Scholar

Development of unigene-derived SSR markers in cowpea (Vigna unguiculata) and their transferability to other Vigna species
![The newly developed genomic-SSR markers uncover the genetic characteristics and relationships of olive accessions [PeerJ] The newly developed genomic-SSR markers uncover the genetic characteristics and relationships of olive accessions [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2020/8573/1/fig-2-1x.jpg)
The newly developed genomic-SSR markers uncover the genetic characteristics and relationships of olive accessions [PeerJ]
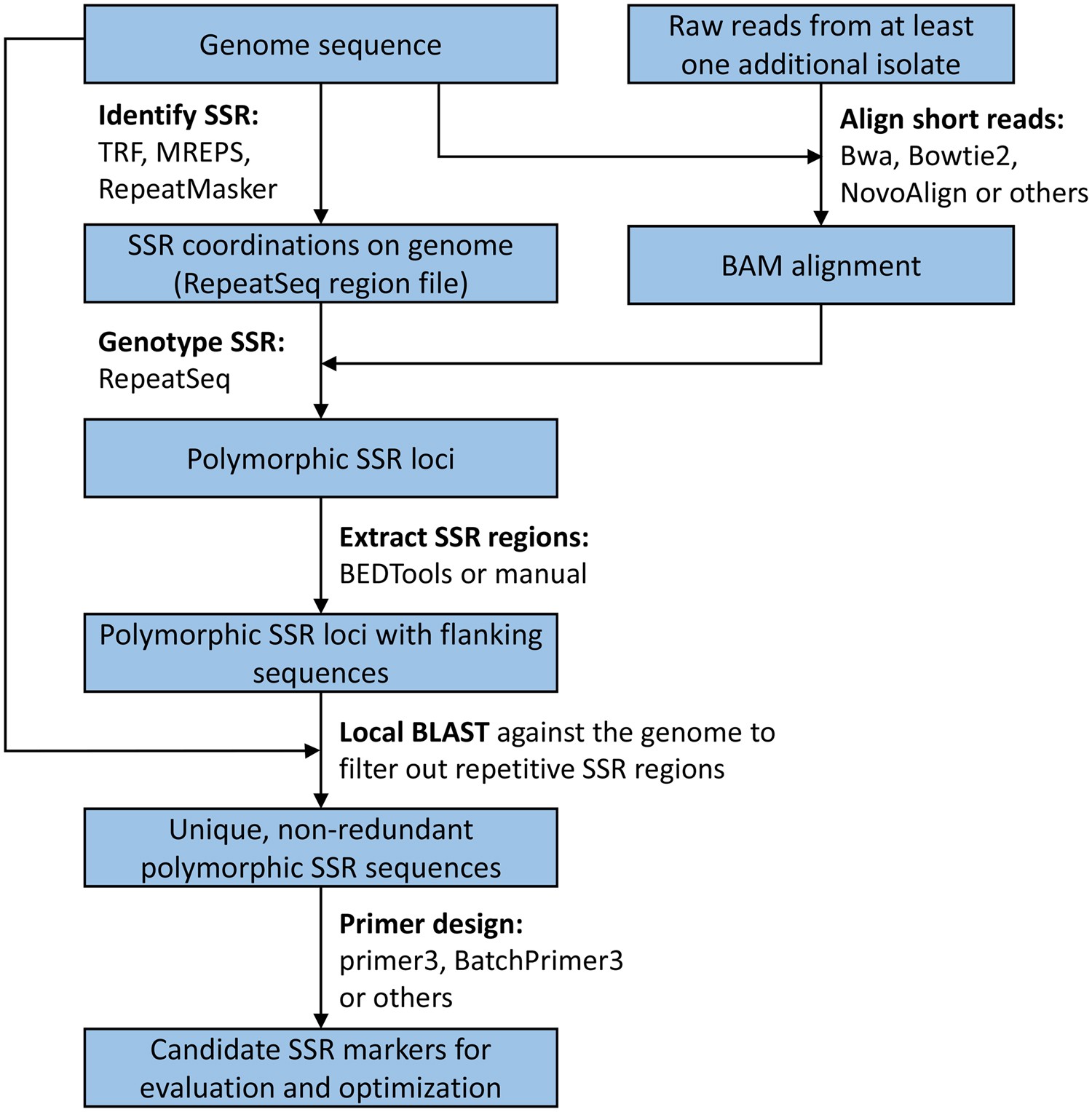
New microsatellite markers for population studies of Phytophthora cinnamomi, an important global pathogen | Scientific Reports
Development of novel EST-SSR markers for ploidy identification based on de novo transcriptome assembly for Misgurnus anguillicaudatus | PLOS ONE
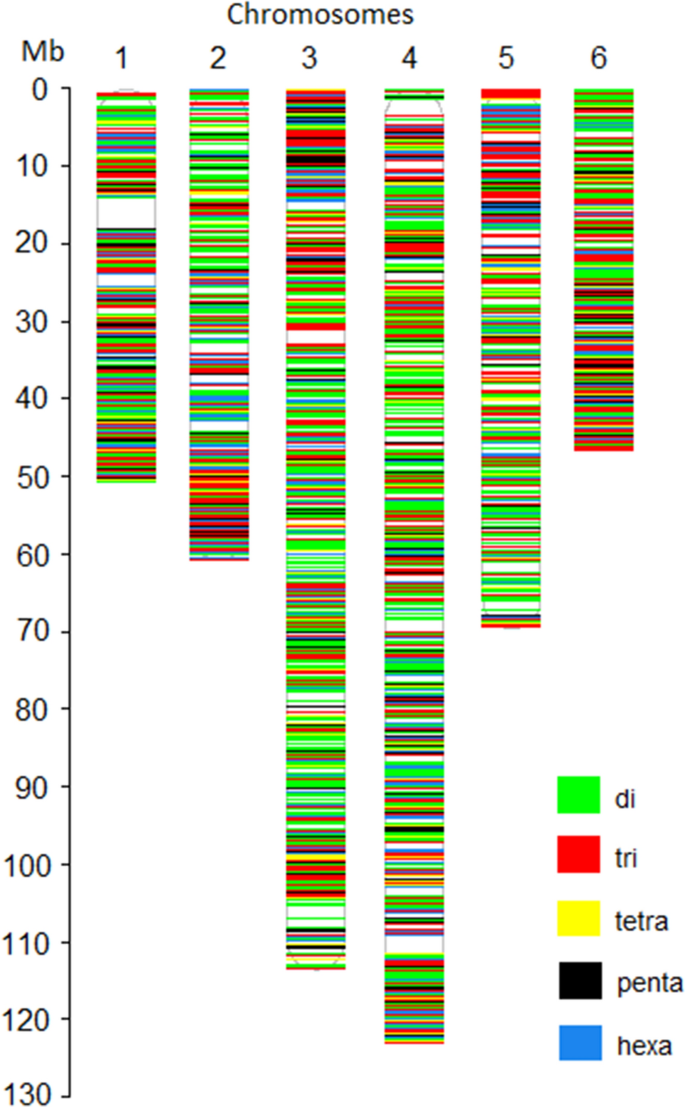
Genome-wide simple sequence repeats (SSR) markers discovered from whole-genome sequence comparisons of multiple spinach accessions | Scientific Reports
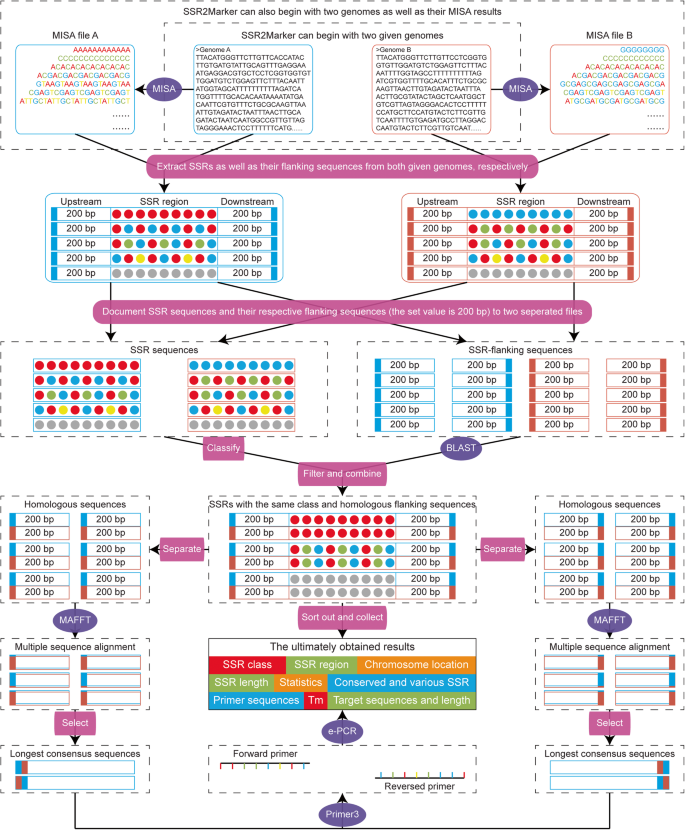
SSR2Marker: an integrated pipeline for identification of SSR markers within any two given genome-scale sequences | Molecular Horticulture | Full Text
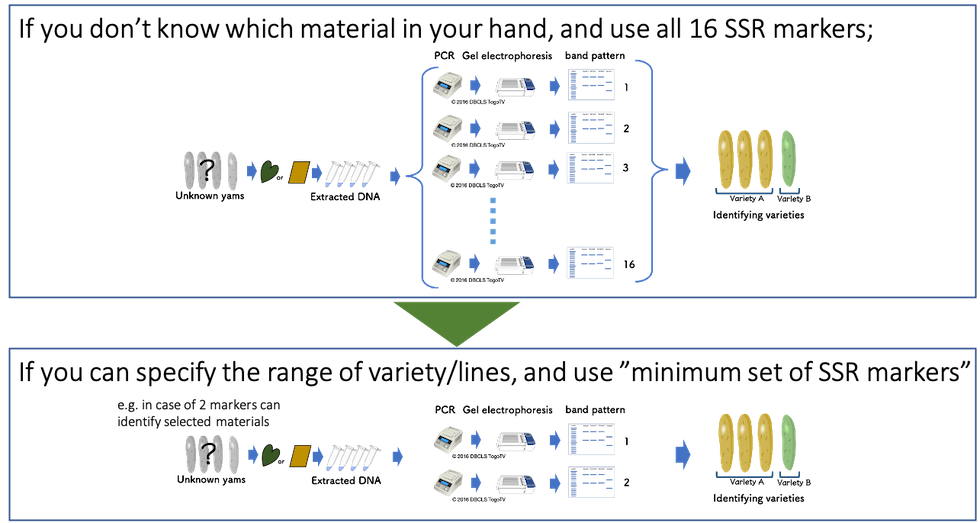
Yam Variety Identification Toolkit:Step 1 | Japan International Research Center for Agricultural Sciences | JIRCAS