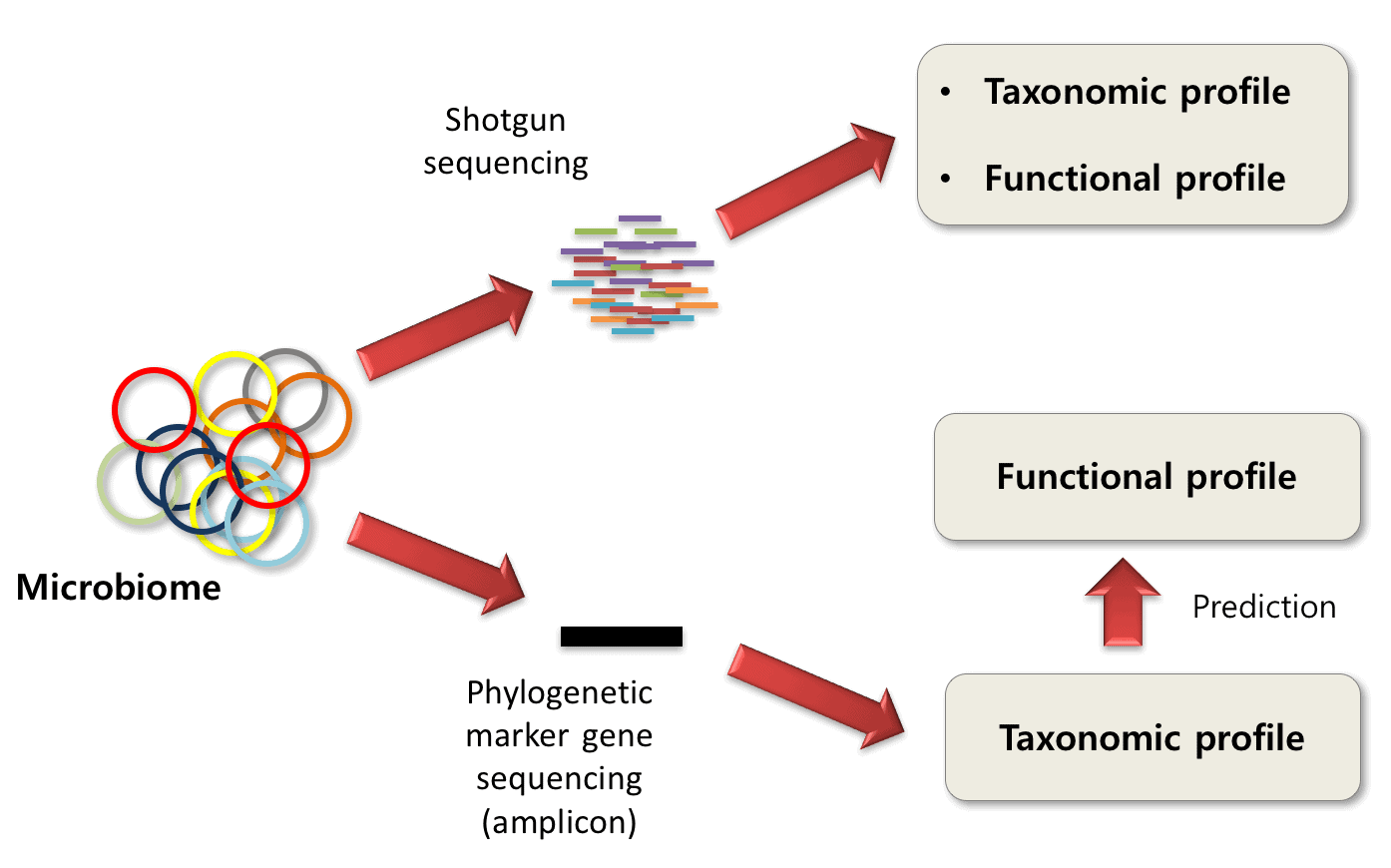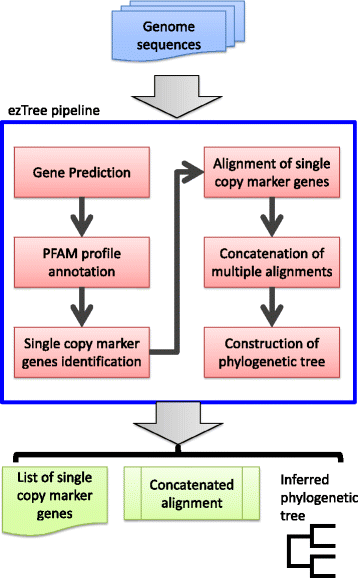
ezTree: an automated pipeline for identifying phylogenetic marker genes and inferring evolutionary relationships among uncultivated prokaryotic draft genomes | BMC Genomics | Full Text
The Methanol Dehydrogenase Gene, mxaF, as a Functional and Phylogenetic Marker for Proteobacterial Methanotrophs in Natural Environments | PLOS ONE

ALG11 – A new variable DNA marker for sponge phylogeny: Comparison of phylogenetic performances with the 18S rDNA and the COI gene - ScienceDirect
Comparative phylogeny of 36 conserved phylogenetic marker proteins and... | Download Scientific Diagram
![The transferability of microsatellite loci from a homoploid to a polyploid hybrid complex: an example from fine-leaved Festuca species (Poaceae) [PeerJ] The transferability of microsatellite loci from a homoploid to a polyploid hybrid complex: an example from fine-leaved Festuca species (Poaceae) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2020/9227/1/fig-2-1x.jpg)
The transferability of microsatellite loci from a homoploid to a polyploid hybrid complex: an example from fine-leaved Festuca species (Poaceae) [PeerJ]

Development of phylogenetic markers from single-copy nuclear genes for multi locus, species level analyses in the mint family (Lamiaceae) - ScienceDirect

The phylogenetic tree based on three separate markers, COI, Cyt b, and... | Download Scientific Diagram

The influence of molecular markers and methods on inferring the phylogenetic relationships between the representatives of the Arini (parrots, Psittaciformes), determined on the basis of their complete mitochondrial genomes | BMC Ecology
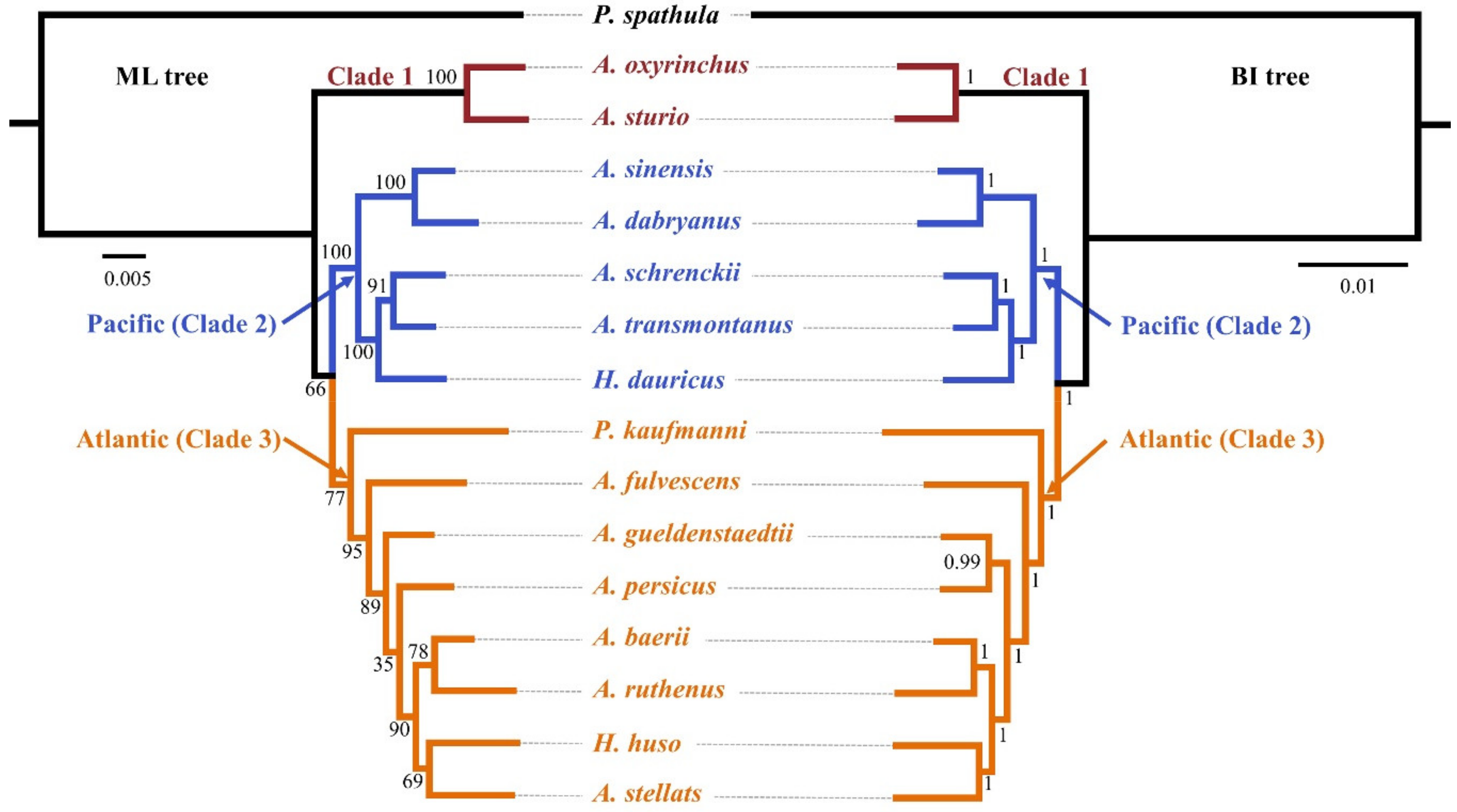
Genes | Free Full-Text | Highly Resolved Phylogenetic Relationships within Order Acipenseriformes According to Novel Nuclear Markers
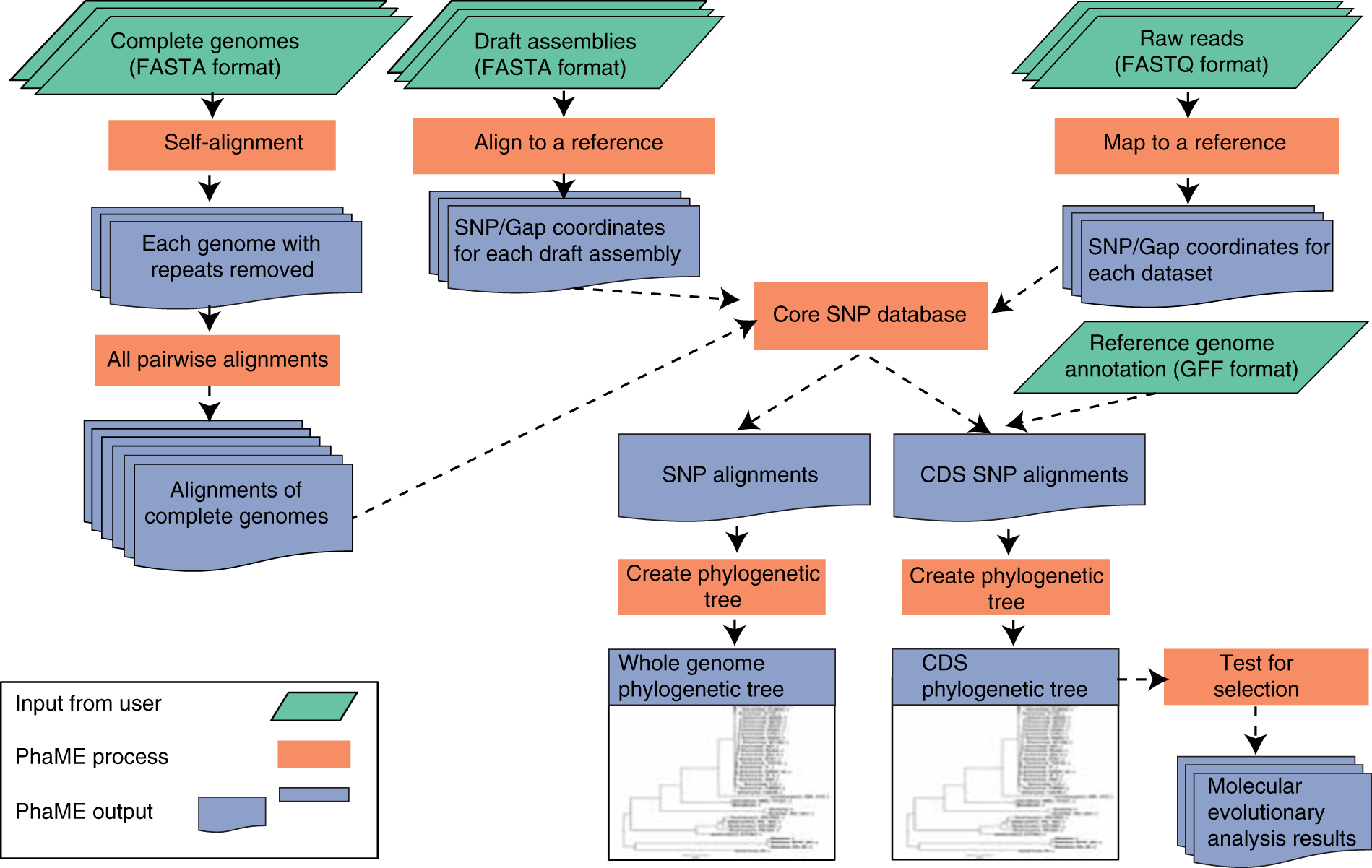
Standardized phylogenetic and molecular evolutionary analysis applied to species across the microbial tree of life | Scientific Reports
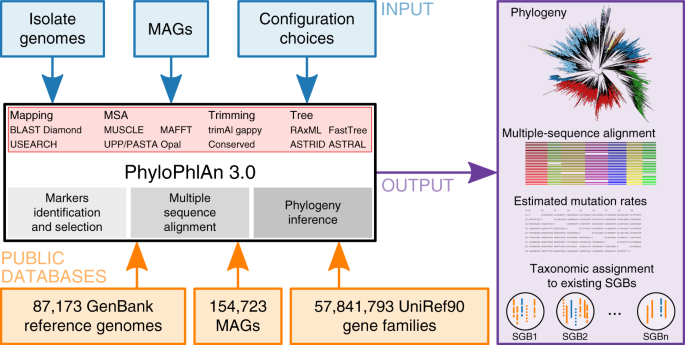
Precise phylogenetic analysis of microbial isolates and genomes from metagenomes using PhyloPhlAn 3.0 | Nature Communications
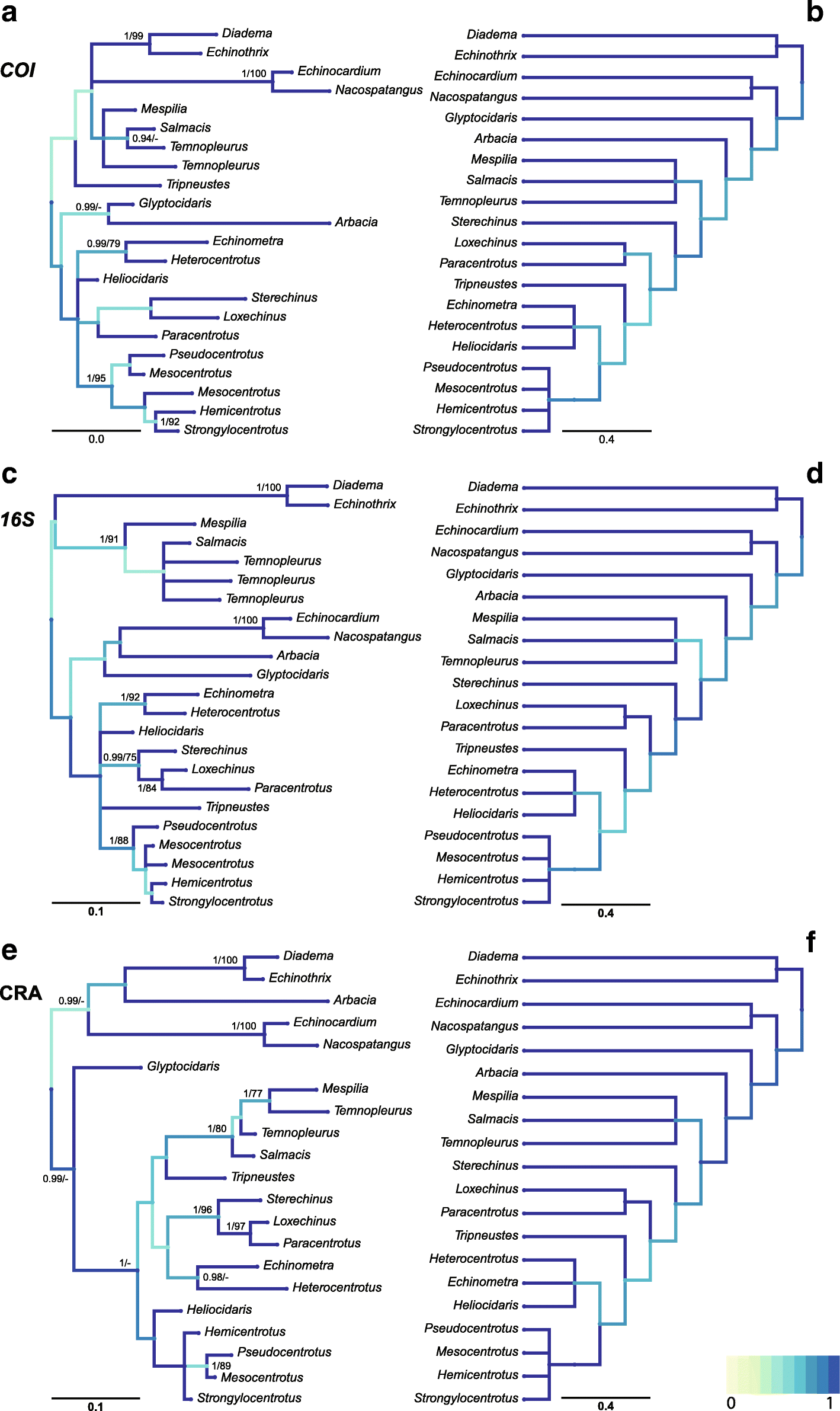
Mind the gap! The mitochondrial control region and its power as a phylogenetic marker in echinoids | BMC Ecology and Evolution | Full Text

Popmarker: Identifying Phylogenetic Markers at the Population Level - Huei-Mien Ke, Chun-Ping Yu, Yu-Ching Liu, Isheng J Tsai, 2017
Identifying genetic markers for a range of phylogenetic utility–From species to family level | PLOS ONE