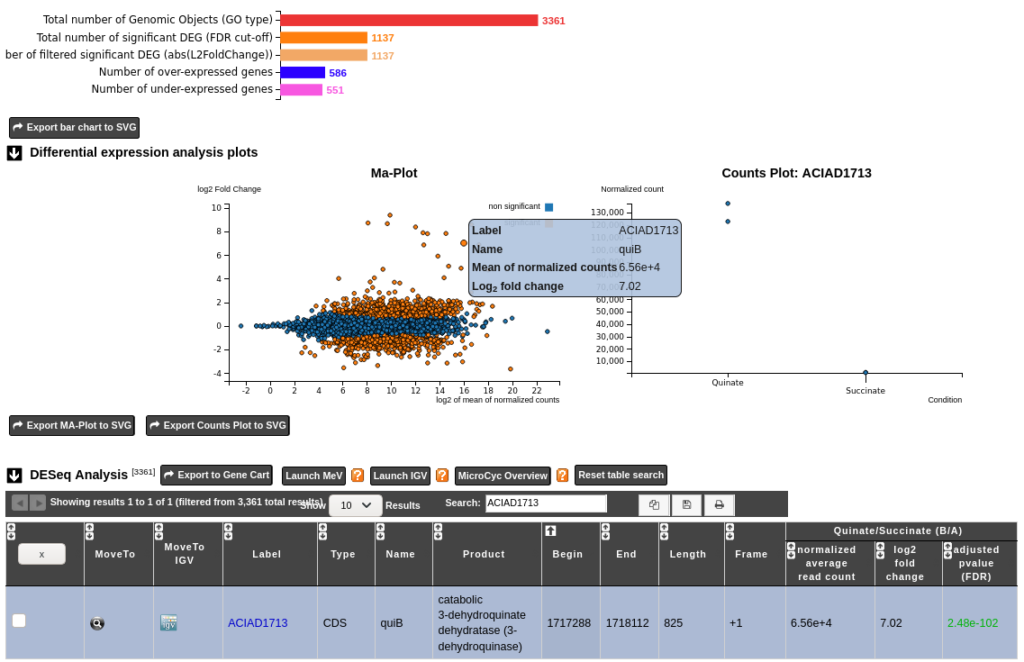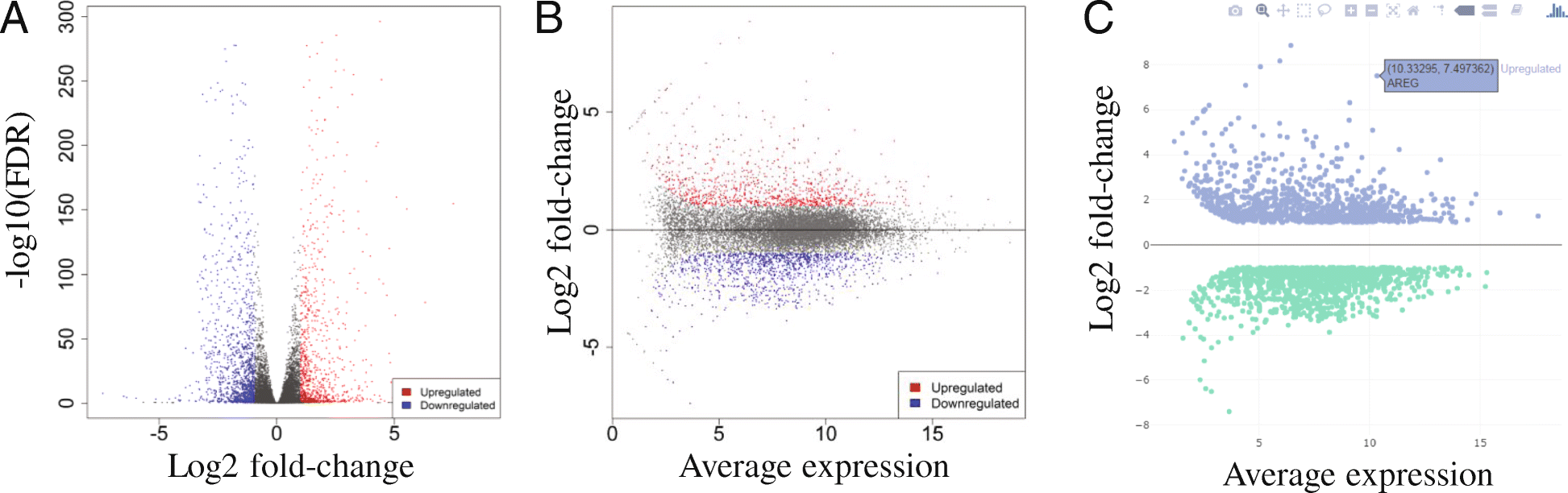
iDEP: an integrated web application for differential expression and pathway analysis of RNA-Seq data | BMC Bioinformatics | Full Text

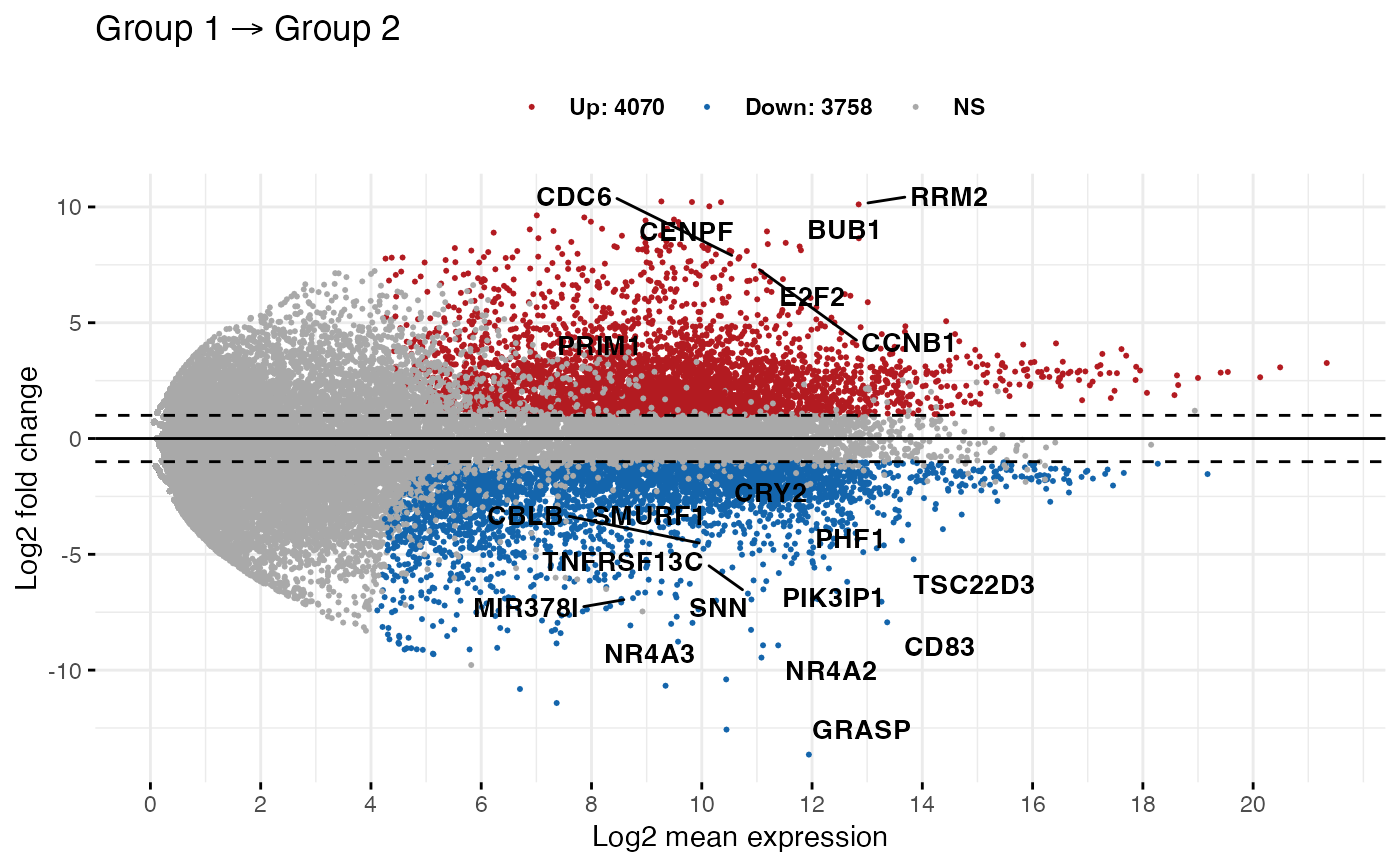
Genome accessibility dynamics in response to phosphate limitation is controlled by the PHR1 family of transcription factors in Arabidopsis | PNAS
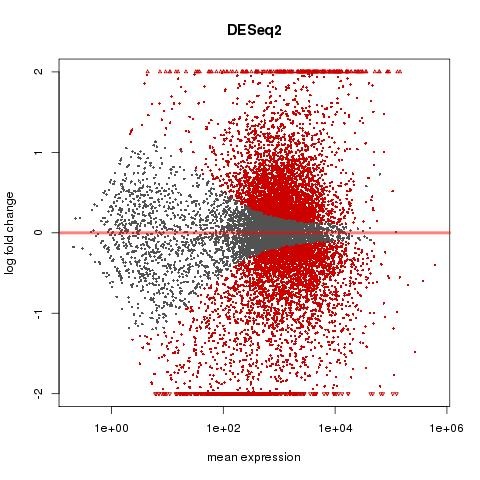
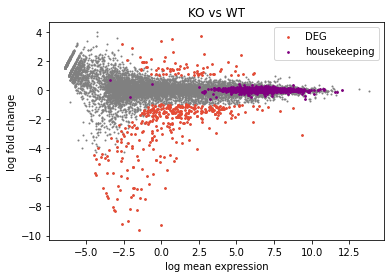
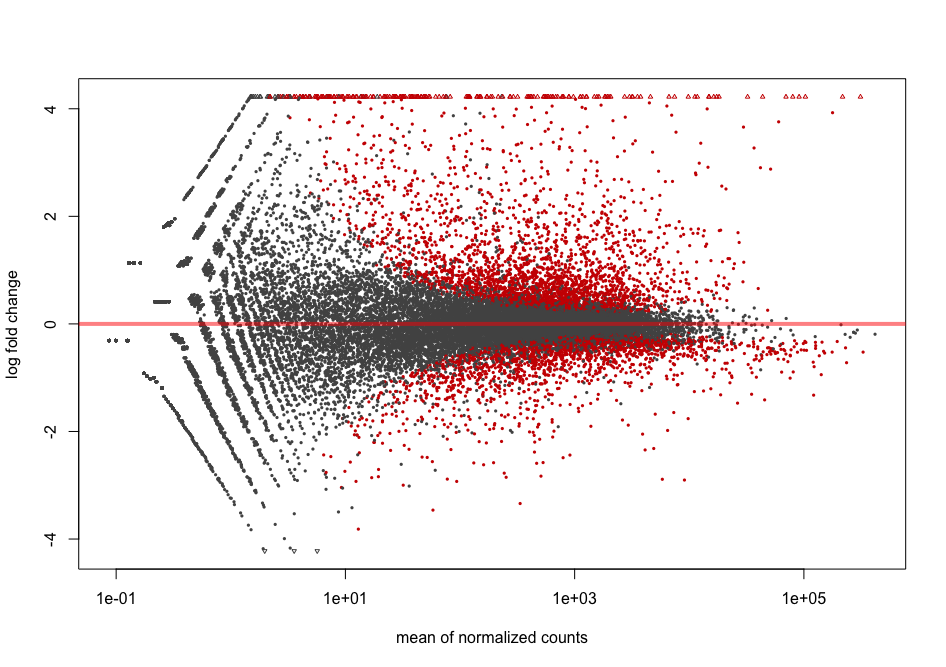
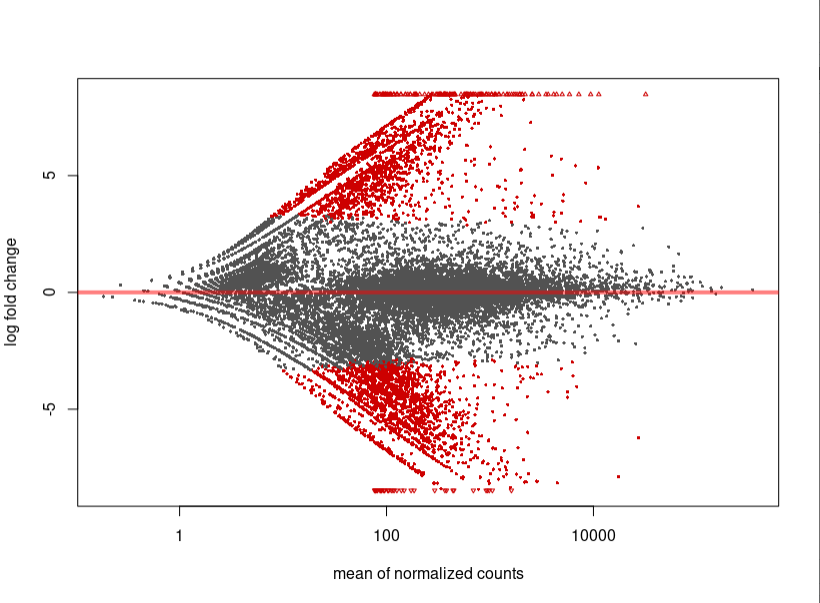
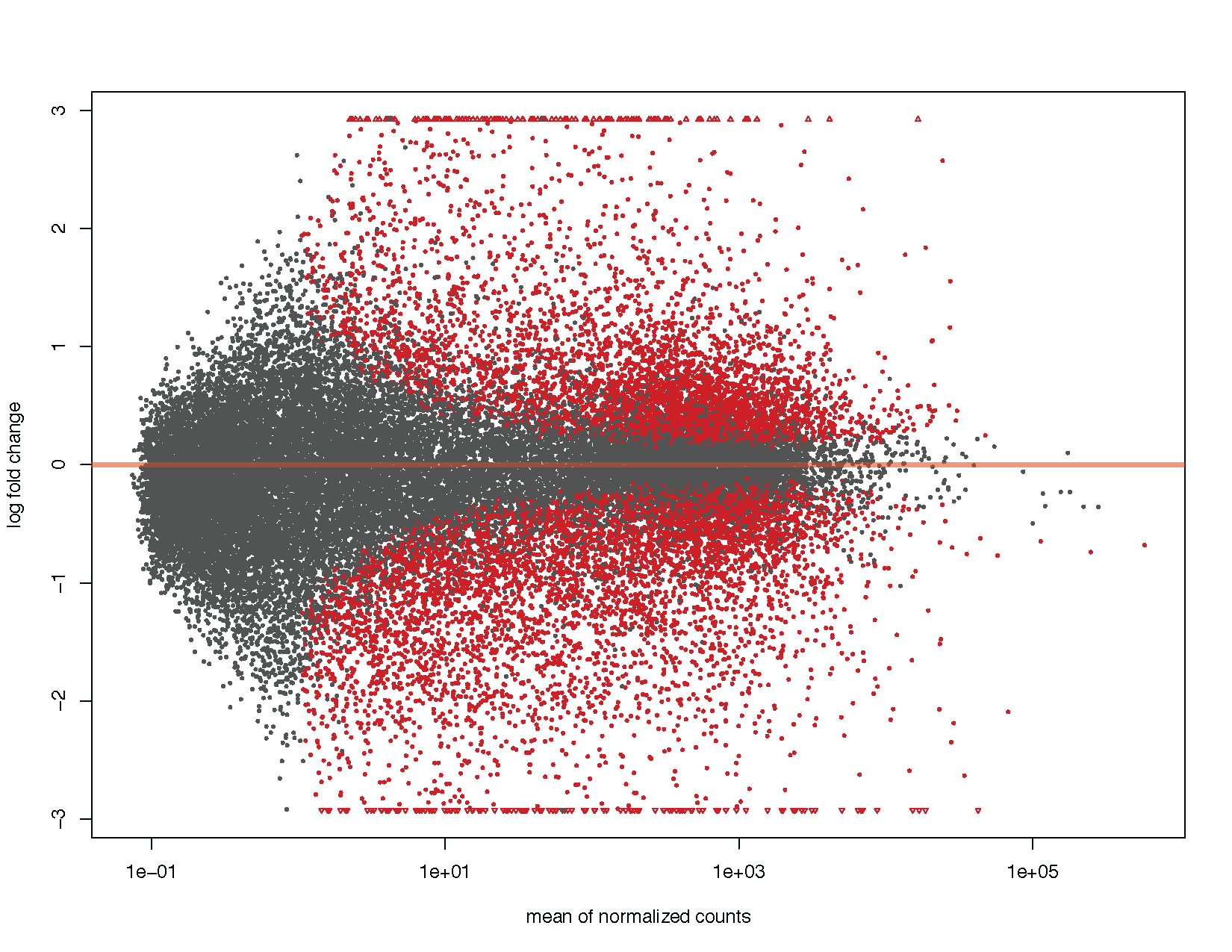
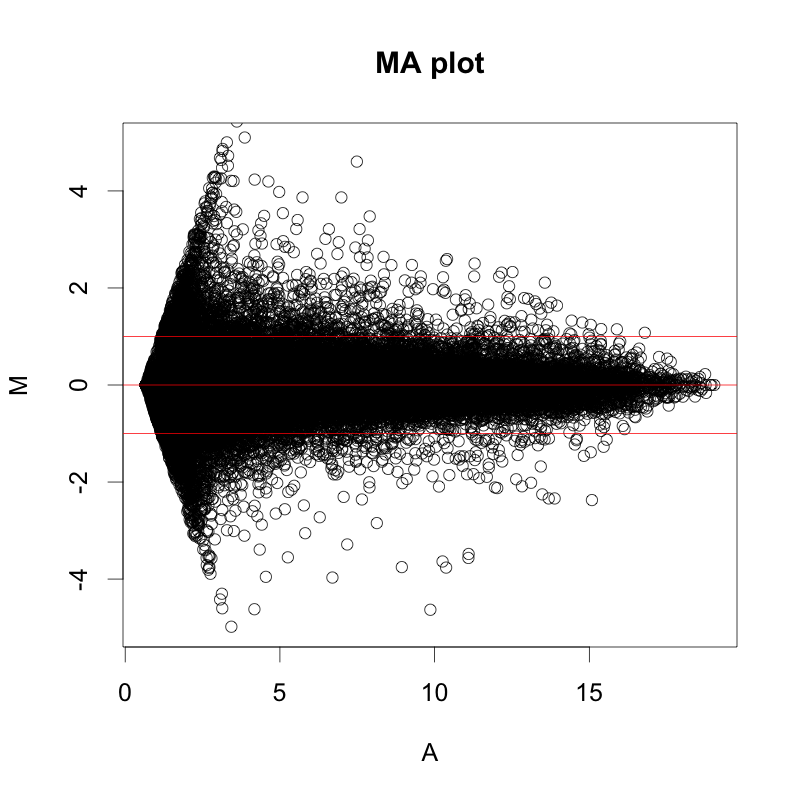
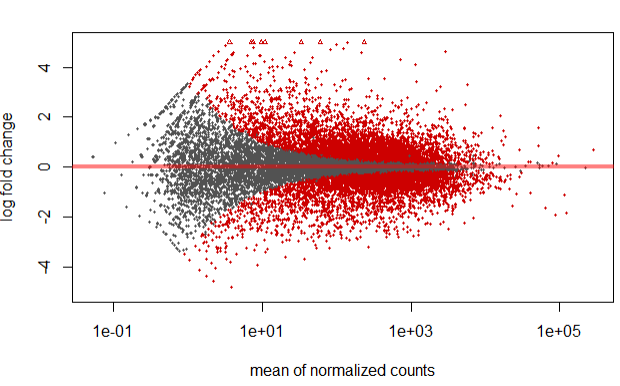
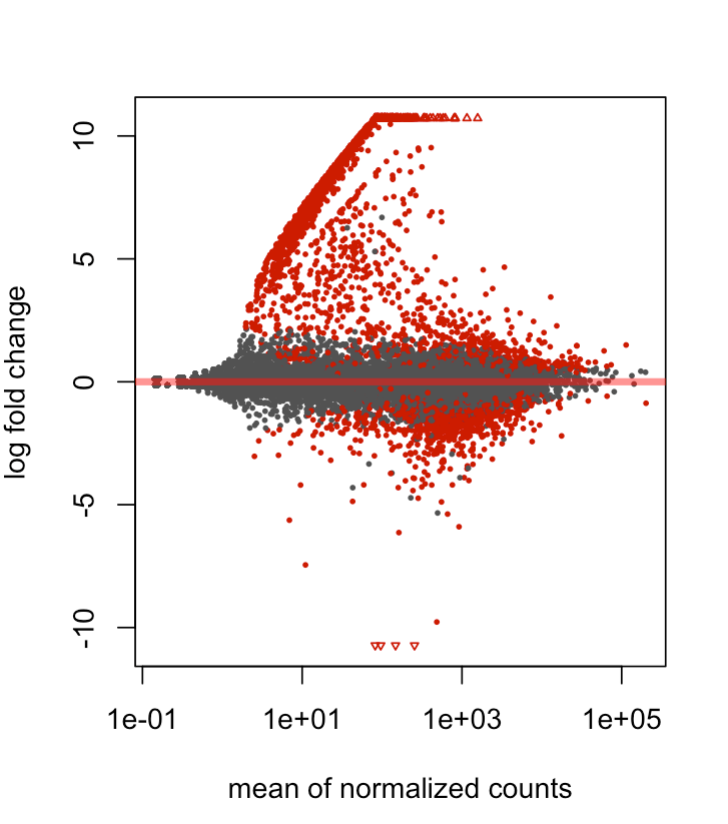
Plotting DESEQ2 Results - Bioinformatics Team (BioITeam) at the University of Texas - UT Austin Wikis