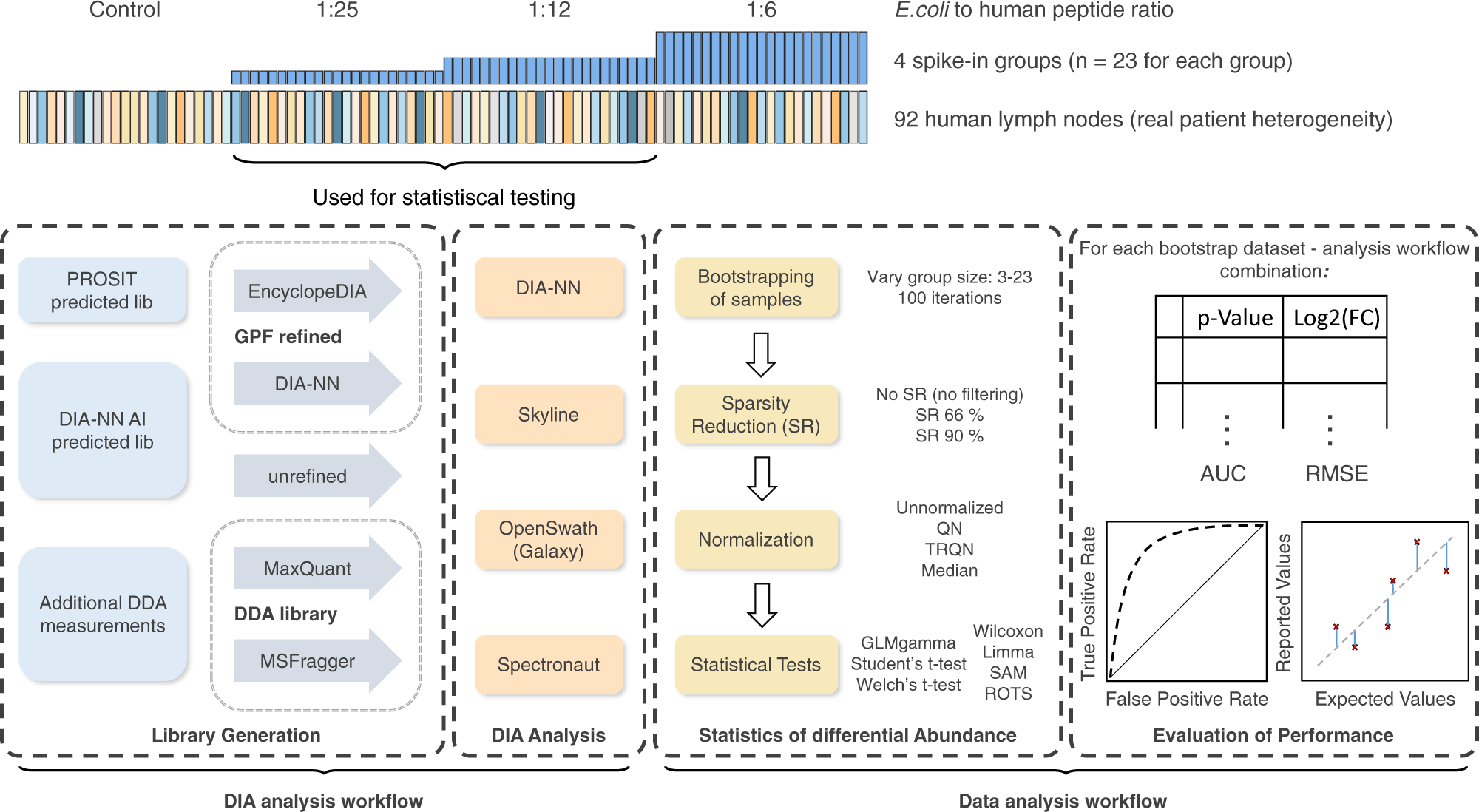
Benchmarking of analysis strategies for data-independent acquisition proteomics using a large-scale dataset comprising inter-patient heterogeneity | Nature Communications

Comparison of Full-Scan, Data-Dependent, and Data-Independent Acquisition Modes in Liquid Chromatography–Mass Spectrometry Based Untargeted Metabolomics | Analytical Chemistry
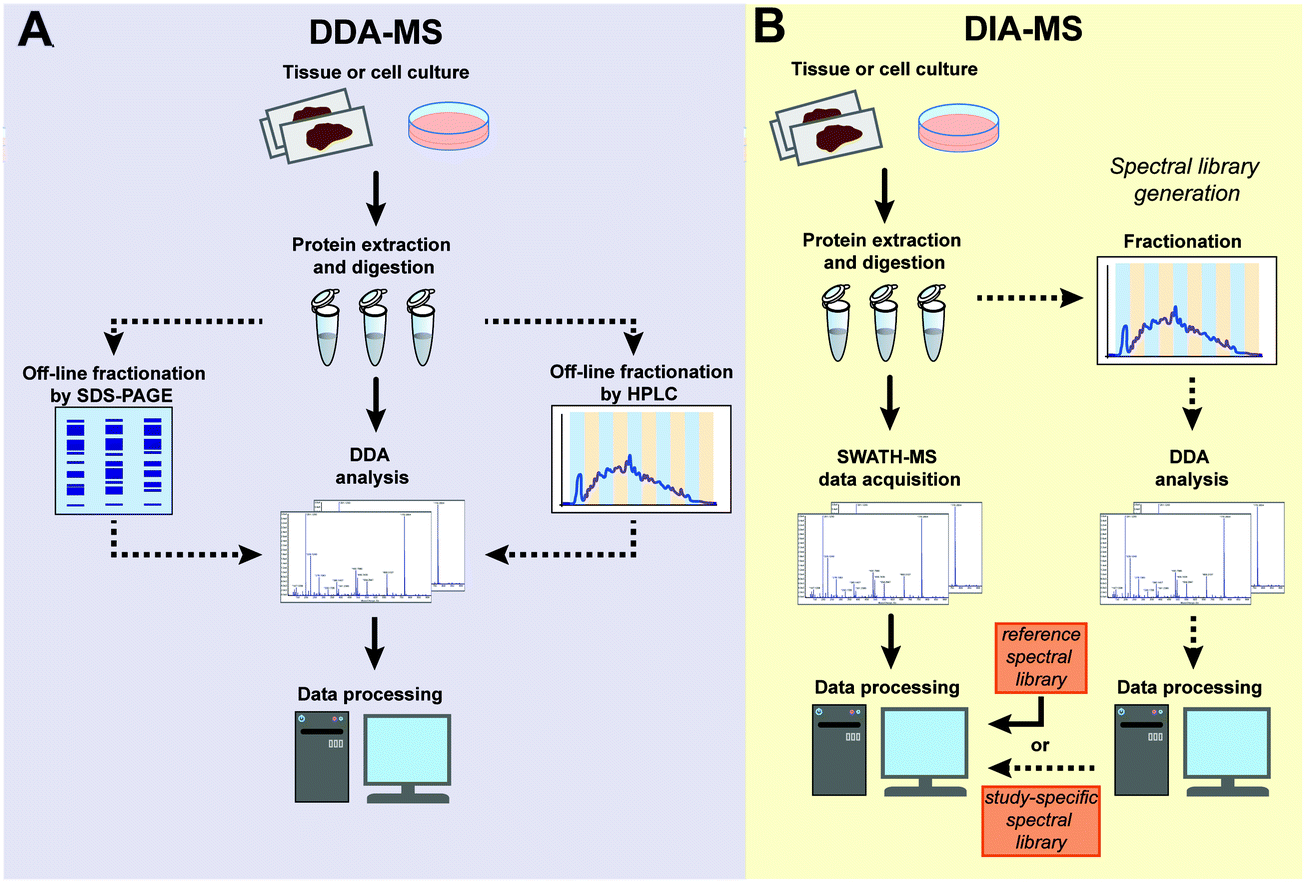
Data-independent acquisition mass spectrometry (DIA-MS) for proteomic applications in oncology - Molecular Omics (RSC Publishing) DOI:10.1039/D0MO00072H
![PDF] Technical advances in proteomics: new developments in data-independent acquisition | Semantic Scholar PDF] Technical advances in proteomics: new developments in data-independent acquisition | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/bbb8bb9323a00f5eefb5d43e3dbb92f9f42e031f/3-Figure1-1.png)
PDF] Technical advances in proteomics: new developments in data-independent acquisition | Semantic Scholar

Comparison of Full-Scan, Data-Dependent, and Data-Independent Acquisition Modes in Liquid Chromatography–Mass Spectrometry Based Untargeted Metabolomics | Analytical Chemistry

Data-independent acquisition method for ubiquitinome analysis reveals regulation of circadian biology | Nature Communications

Data‐independent acquisition‐based SWATH‐MS for quantitative proteomics: a tutorial | Molecular Systems Biology
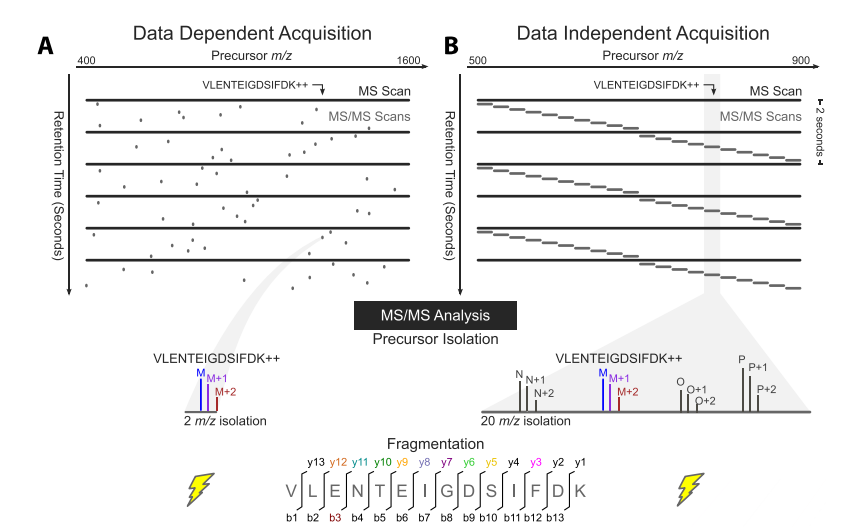
Data Dependent Acquisition and Data Independent Acquisition Mass Spectrometry – Creative Proteomics Blog

The use of hybrid data-dependent and -independent acquisition spectral libraries empower dual-proteome profiling | bioRxiv
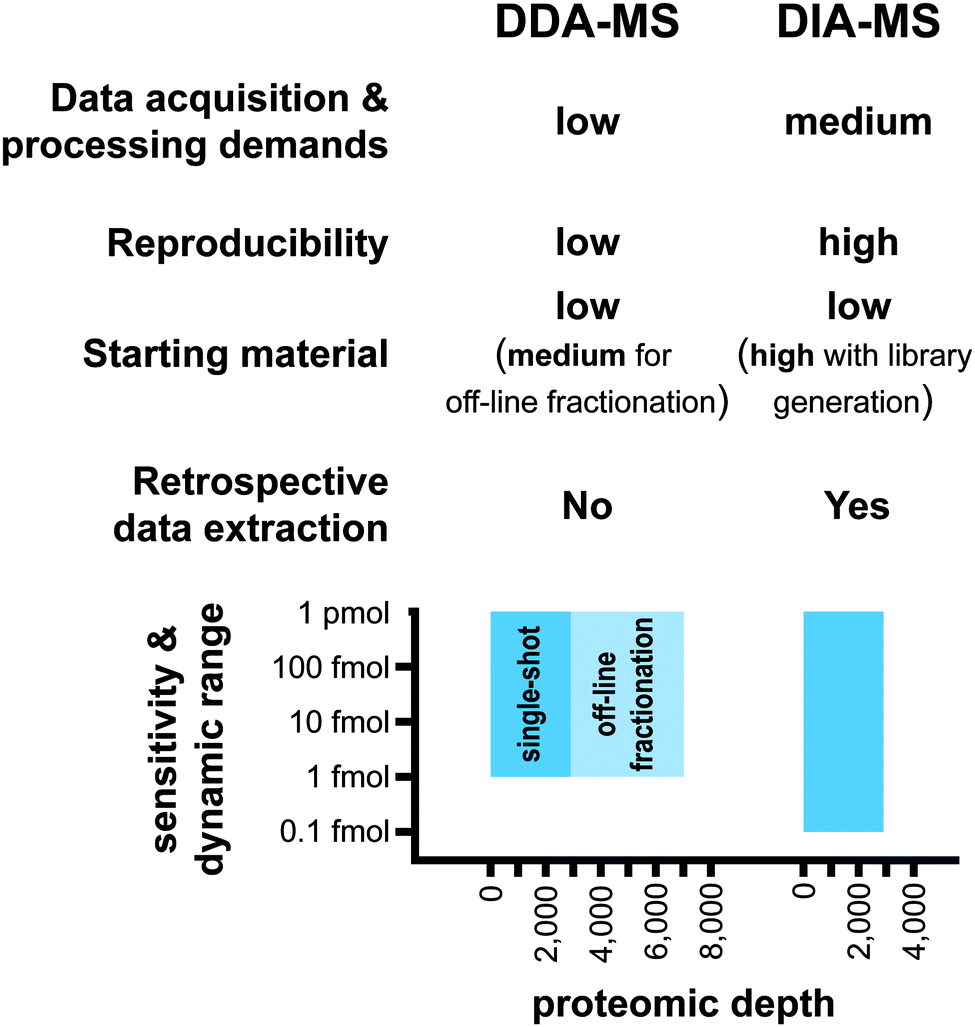
Data-independent acquisition mass spectrometry (DIA-MS) for proteomic applications in oncology - Molecular Omics (RSC Publishing) DOI:10.1039/D0MO00072H

Technical comparison of DDA and different types of DIA in a biological... | Download Scientific Diagram

Advances in data‐independent acquisition mass spectrometry towards comprehensive digital proteome landscape - Kitata - Mass Spectrometry Reviews - Wiley Online Library
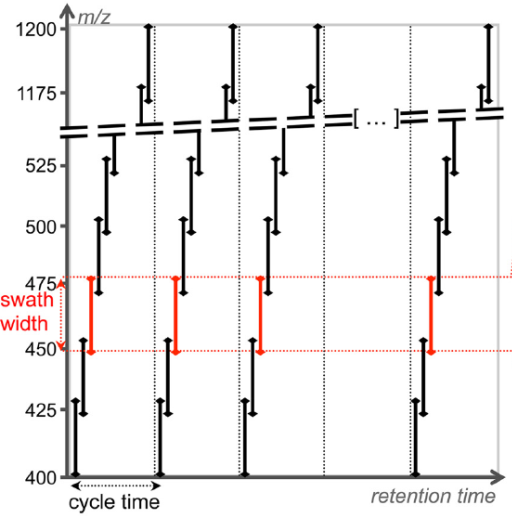
Data Dependent Acquisition and Data Independent Acquisition Mass Spectrometry – Creative Proteomics Blog