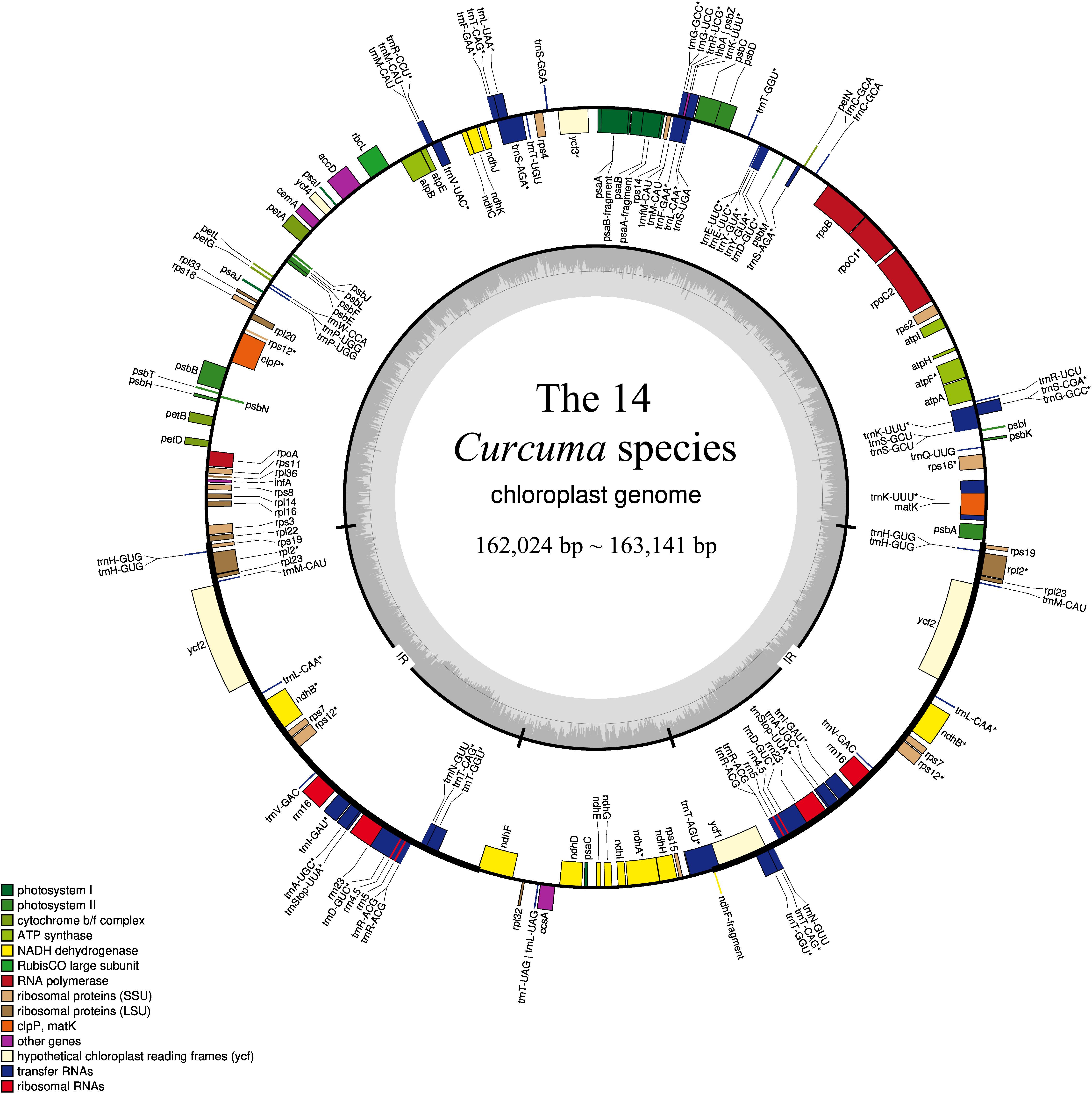
Comparative Chloroplast Genomes of Sorghum Species: Sequence Divergence and Phylogenetic Relationships
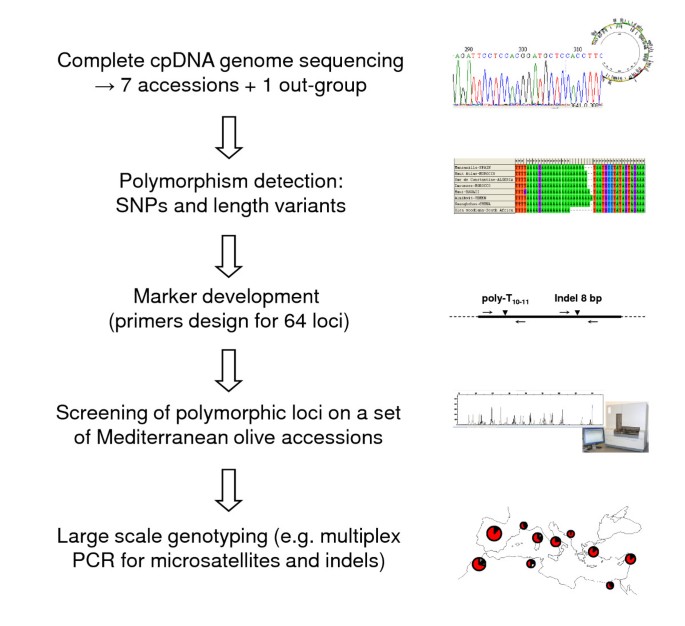
Genomic profiling of plastid DNA variation in the Mediterranean olive tree | BMC Plant Biology | Full Text

Identifying the North American Plum Species Phylogenetic Signal Using Nuclear, Mitochondrial, and Chloroplast DNA Markers in: Journal of the American Society for Horticultural Science Volume 141 Issue 6 (2016)
Identifying the North American Plum Species Phylogenetic Signal Using Nuclear, Mitochondrial, and Chloroplast DNA Markers

The complete chloroplast genomes of three Hamamelidaceae species: Comparative and phylogenetic analyses - Wang - 2022 - Ecology and Evolution - Wiley Online Library

Horticulturae | Free Full-Text | Comparative Analyses of 18 Complete Chloroplast Genomes from Eleven Mangifera Species (Anacardiaceae): Sequence Characteristics and Phylogenomics

Comparison of genetic differentiation at cpDNA markers between Quercus... | Download Scientific Diagram

Comparison of Four Complete Chloroplast Genomes of Medicinal and Ornamental Meconopsis Species: Genome Organization and Species Discrimination | Scientific Reports

Chloroplast DNA: A Promising Source of Information for Plant Phylogeny and Traceability | SciTechnol
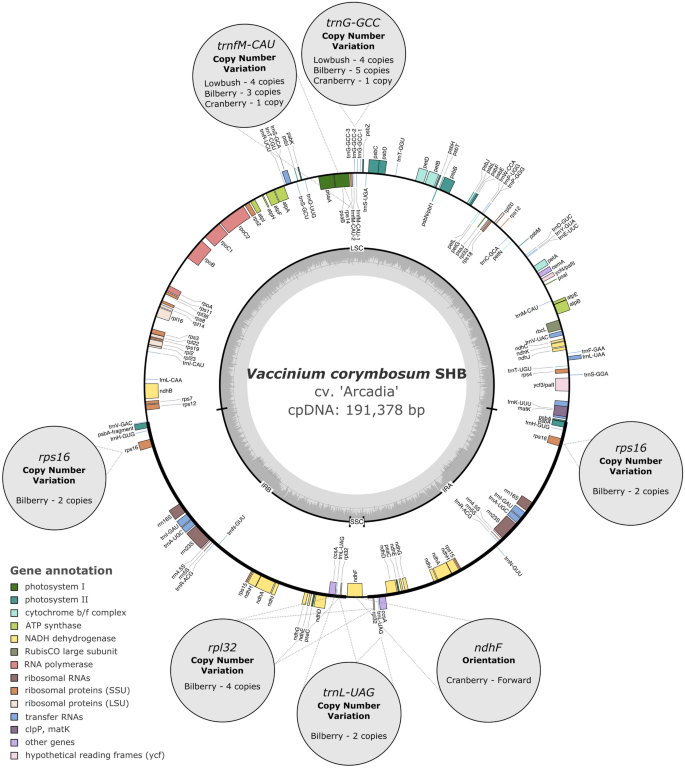
Chloroplast genome assemblies and comparative analyses of commercially important Vaccinium berry crops | Scientific Reports
Genetic diversity and relationship between cultivated, weedy and wild rye species as revealed by chloroplast and mitochondrial DNA non-coding regions analysis | PLOS ONE
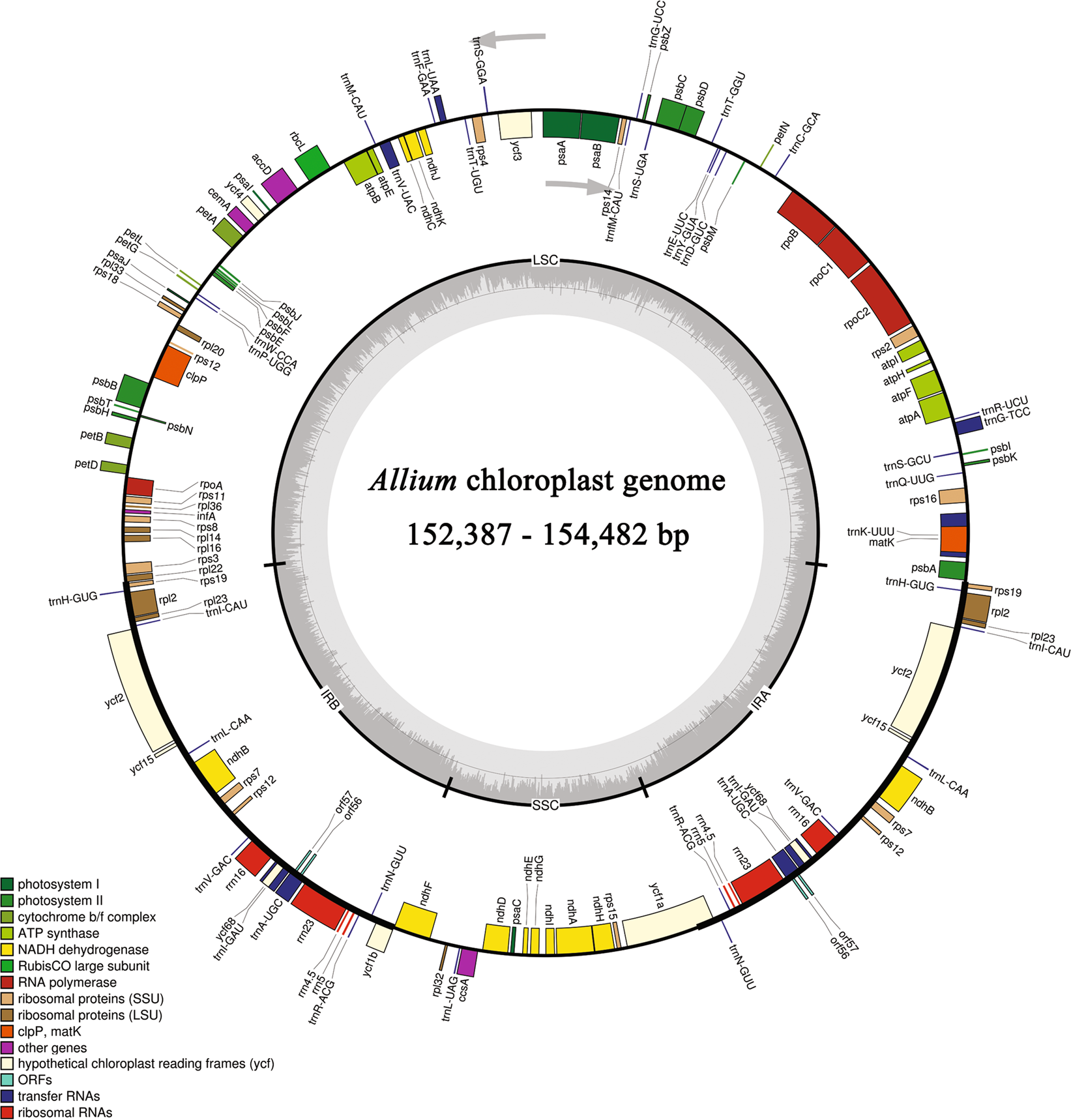
Complete chloroplast genome sequences of four Allium species: comparative and phylogenetic analyses | Scientific Reports
![PDF] Evaluation of chloroplast DNA markers for distinguishing Phalaenopsis species | Semantic Scholar PDF] Evaluation of chloroplast DNA markers for distinguishing Phalaenopsis species | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/fa8fb7d7625c2b5d1bf02b05b1d533640f2b53c1/4-Table3-1.png)
PDF] Evaluation of chloroplast DNA markers for distinguishing Phalaenopsis species | Semantic Scholar
![Comparison of the effectiveness of ISJ and SSR markers and detection of outlier loci in conservation genetics of Pulsatilla patens populations [PeerJ] Comparison of the effectiveness of ISJ and SSR markers and detection of outlier loci in conservation genetics of Pulsatilla patens populations [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2016/2504/1/fig-3-full.png)
Comparison of the effectiveness of ISJ and SSR markers and detection of outlier loci in conservation genetics of Pulsatilla patens populations [PeerJ]

Molecular Genetic Diversity and Phylogenetic Analyses of Punica granatum L. Populations Revealed by ISSR Markers and Chloroplast (cpDNA) trnL-F Region | SpringerLink

SciELO - Brasil - Microsatellite markers: what they mean and why they are so useful Microsatellite markers: what they mean and why they are so useful