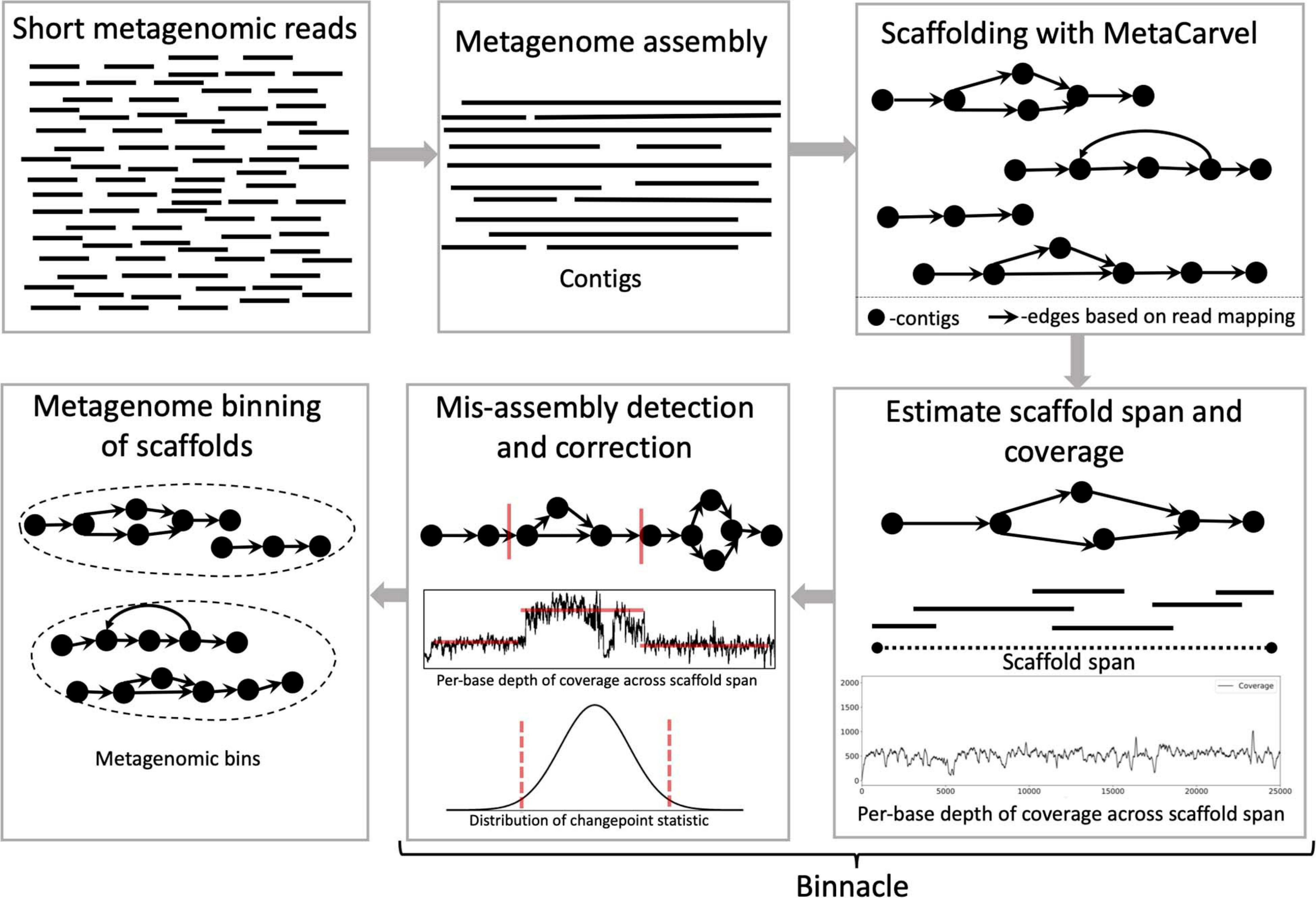
Scaffolding and completing genome assemblies in real-time with nanopore sequencing | Nature Communications

The changing face of genome assemblies: Guidance on achieving high‐quality reference genomes - Whibley - 2021 - Molecular Ecology Resources - Wiley Online Library
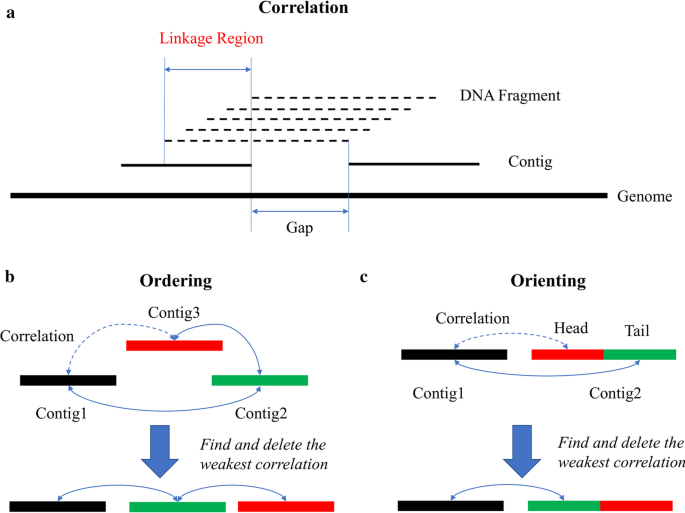
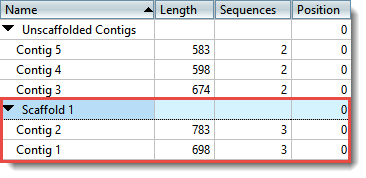
SLR-superscaffolder: a de novo scaffolding tool for synthetic long reads using a top-to-bottom scheme | BMC Bioinformatics | Full Text
Plants | Free Full-Text | Reference-Guided De Novo Genome Assembly to Dissect a QTL Region for Submergence Tolerance Derived from Ciherang-Sub1

Types of assembled contigs and alignments to REF contigs. a A contig... | Download Scientific Diagram

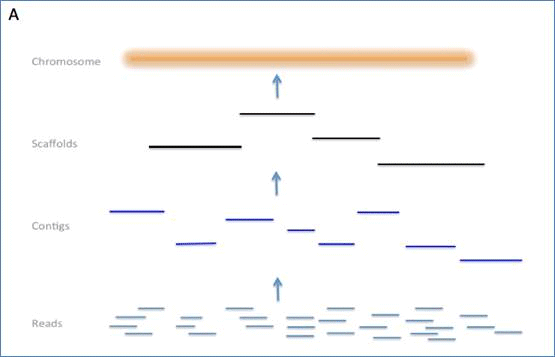
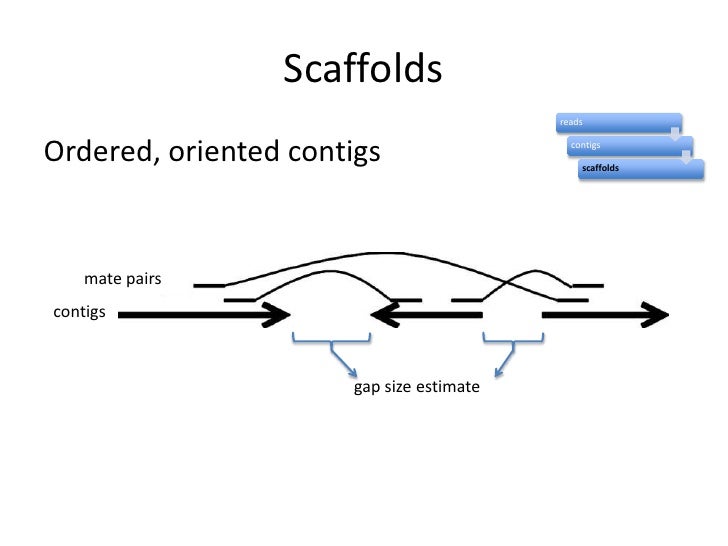
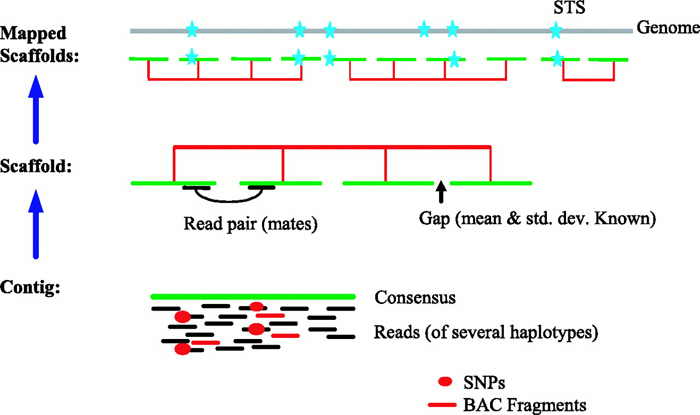
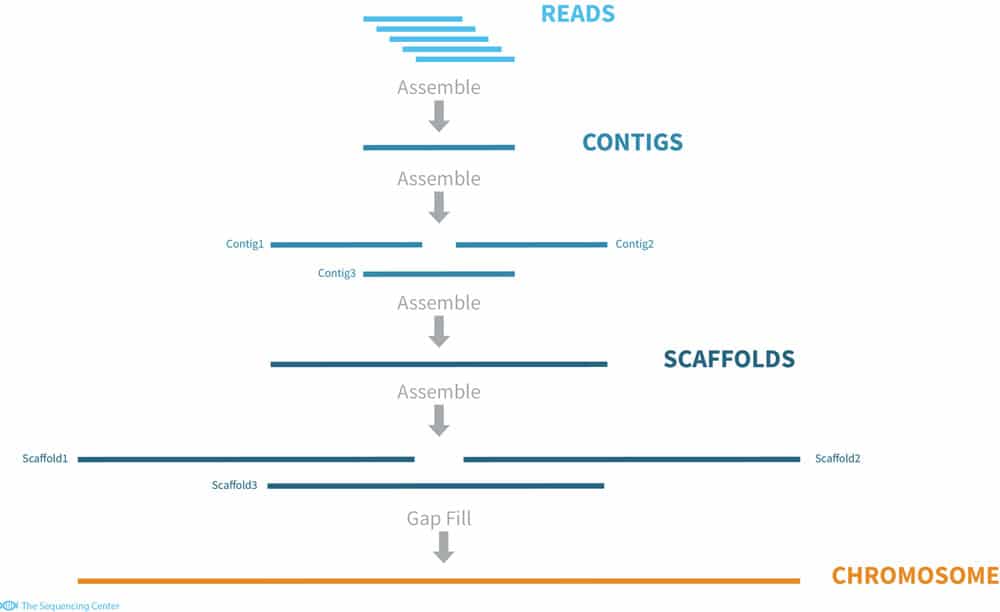
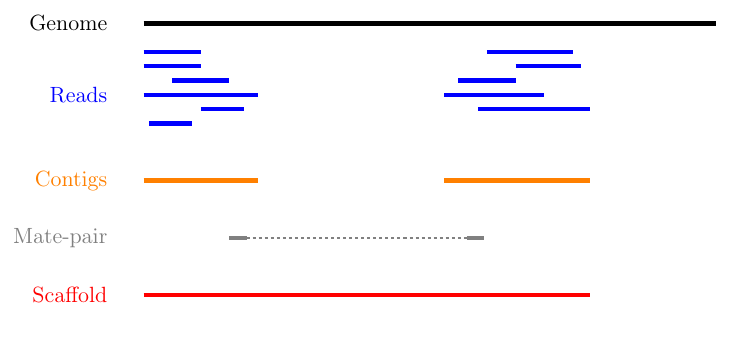
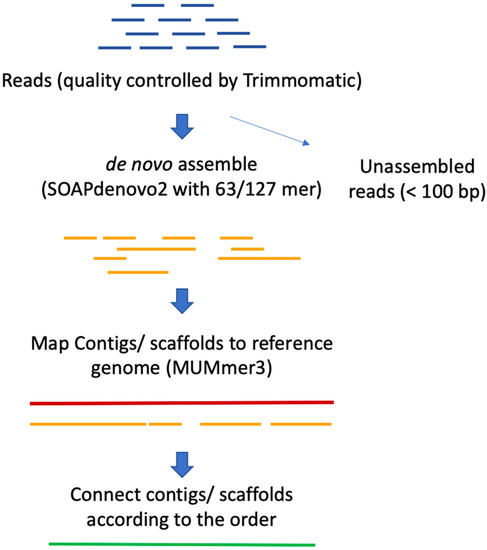
Schematic representation of the method used to assemble Illumina reads into contigs, and contigs into scaffolds.