A proteomics approach to identify targets of the ubiquitin-like molecule Urm1 in Drosophila melanogaster | PLOS ONE

A Magnetic Bead–Based Ligand Binding Assay to Facilitate Human Kynurenine 3-Monooxygenase Drug Discovery - Kris Wilson, Damian J. Mole, Natalie Z. M. Homer, John P. Iredale, Manfred Auer, Scott P. Webster, 2015

Functional proteomics protocol for the identification of interaction partners in Tetrahymena thermophila - ScienceDirect
Supplemental Information The mRNA-Bound Proteome and Its Global Occupancy Profile on Protein-Coding Transcripts

A proteomics protocol to identify stimulation-induced binding partners dependent on a specific gene in mammalian cells - ScienceDirect

Design and Characterization of a New pVII Combinatorial Phage Display Peptide Library for Protease Substrate Mining Using Factor VII Activating Protease (FSAP) as Model - Kara - 2020 - ChemBioChem - Wiley Online Library
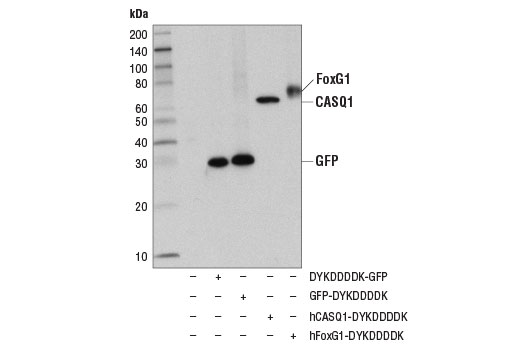
DYKDDDDK Tag (D6W5B) Rabbit mAb (Binds to same epitope as Sigma's Anti-FLAG® M2 Antibody) | Cell Signaling Technology
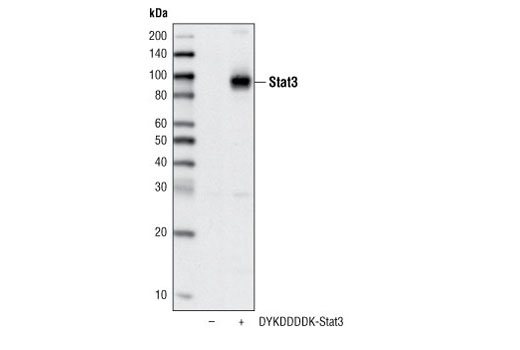
DYKDDDDK Tag Antibody (Binds to same epitope as Sigma's Anti-FLAG® M2 Antibody) | Cell Signaling Technology